DNA-Encoded Chemical Library Screening Is a Great Tool for Identifying Novel Binders Suitable for Bispecific Degraders
The Value of DEL for PROTACs and Other Bispecifics
Proteolysis targeting chimeras (PROTACs) are bispecific molecules that induce protein degradation of a specific target. These compounds have been a hot area of chemistry and pharma since they were discovered around 20 years ago.
Because of their structure, PROTACs have long been considered the perfect application for DNA-Encoded Library (DEL) screening. PROTACS have modular structures, consisting of a target binding moiety, a linker, and an engager with an E3 ligase. What makes DEL perfect for creating this bispecific structure is that in the library, the linkage point is built in already, connecting the discovered compound to its DNA label. In fact, you even know that the linker is tolerated in the binding event because it was there when you discovered the interaction. So all you need to do is replace the DNA with an appropriate E3 ligase engager and your discovery is ready to go.
Discovery of Bispecific Estrogen Receptor α Degraders With Novel Binders
To demonstrate this concept, our team built a bispecific degrader (PROTAC) of estrogen receptor α (ERα) with the goal of improving upon current modulators of ERα signaling (aromatase inhibitors/SERMs/SERDs) that are used to treat ER+ breast cancer. (For the full details, see our article, Bispecific Estrogen Receptor α Degraders Incorporating Novel Binders Identified Using DNA-Encoded Chemical Library Screening just out in the Journal of Medicinal Chemistry.)
We screened around 120 billion DNA-encoded molecules against wild type ERα andthreegain-of-function (GOF) mutants conferring partial resistance to existing endocrine therapies (Fig. 1a). Once enriched small molecules were identified via their DNA sequences, the small molecule binders were quickly leveraged in a parallel click chemistry array producing PROTACs for ERα with a variety of E3 binders (Fig. 1b).
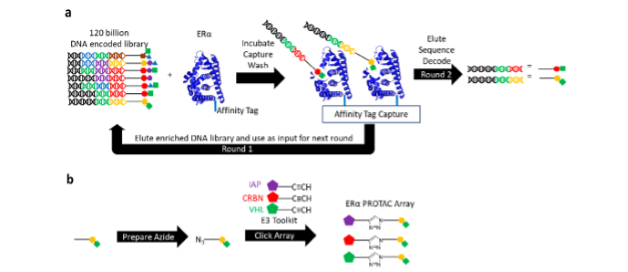
Figure 1. Schematic representation for DNA-Encoded Chemical Library (DECL) screening and PROTAC array platforms. (a) A DECL (combinatorial synthesized small molecules tethered to DNA tags) is incubated with ERα (target protein, affinity selected and washed to remove non-binders). Binding and affinity capture is repeated using the eluate from Round 1 as input to further enrich the library output. The DNA tags of the round 2 selection are sequenced to identify the small molecule binders. (b) Small molecule binders linked to Azide variants are prepared and reacted in a parallel click chemistry library of E3 binder alkynes to obtain an array of PROTACs for ERα.
Reprinted with permission from Disch J.S. et al., Journal of Medicinal Chemistry 2021 64 (8), 5049-5066, DOI: 10.1021/acs.jmedchem.1c00127. Copyright © (2021) American Chemical Society.
Simultaneous Screening Yields Two Potent Off-DNA Compounds Enriched Against the Wild Type and All Three GOF Mutants
Because DEL library selection delivers not just hits, but profiles of hits for multiple targets included in the experiment, it saves significant time. In this case, the primary discovery screen identified two potent compounds that were enriched against the wild type ERα and three of the most common mutants — S463P, Y537S, and D538G (Fig. 2a & b). Additionally, the two compounds were not enriched in the presence of estradiol, indicating that they likely bind the estradiol binding pocket. To confirm the selection output, direct off-DNA analogs of the two potent compounds were synthesized, and their binding affinity was assessed in a fluorescence polarization binding assay (Fig. 2c).
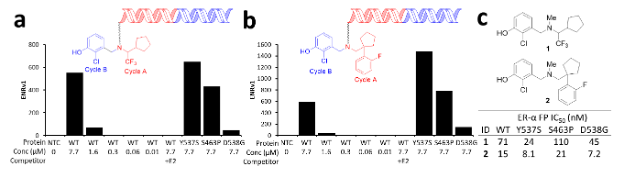
Figure 2. Confirmation of activity of DECL output and off-DNA analogs of two related members. (a, b) Enrichment (ENRv1) profiles of two structurally related members from a 2-cycle library show that they bind to ERα WT and mutants Y537S, S463P, and D538G, but do not bind the matrix (NTC; no target control). Their binding to WT specifically is competed off by the addition of the competitor Estradiol (E2). This suggests that the library members interact at the ligand binding site. (c) Analogous “off-DNA” synthesized exemplars (with linker and DNA truncated to a single methyl group) bind in a fluorescence polarization binding displacement assay. Assay results are an average of duplicate runs.
Reprinted with permission from Disch J.S. et al., Journal of Medicinal Chemistry 2021 64 (8), 5049-5066, DOI: 10.1021/acs.jmedchem.1c00127. Copyright © (2021) American Chemical Society.
Ensuring That the New Compounds Are Binding Antagonistically, Not Agonistically
Even though the goal of this bispecific therapy is to bind and then elicit degradation, it’s important to remember that the small molecule binder could have a cellular effect on its own. In this case the binders we discovered turned out to be agonists when dosed into cells. We were able to convert our binders into antagonists, making small changes to the structure to build a Structure Activity Relationship (SAR) that distinguished antagonism from agonism.
With click chemistry on our most active antagonists, we produced a series of bispecific compounds with variable E3 binding elements and variable linker lengths. Testing for ERα degradation in human breast cancer cells (MCF7) revealed two promising candidate PROTACs: 18 and 21. These two compounds engage VHL as the E3 component; compounds engaging IAP and CRBN did not demonstrate comparable degradation. Both 18 and 21 demonstrated potent binding of wild type ERα and all three GOF mutants (Fig. 3), degraded ERα, led to MCF7 viability defects and downregulated PGR expression.
Despite the low permeability, limited solubility, and high plasma protein binding of 18 and 21, these two PROTACs achieved significant plasma exposures when dosed subcutaneously (Fig. 3f), which warranted evaluation of their ERα degradation activities in vivo.
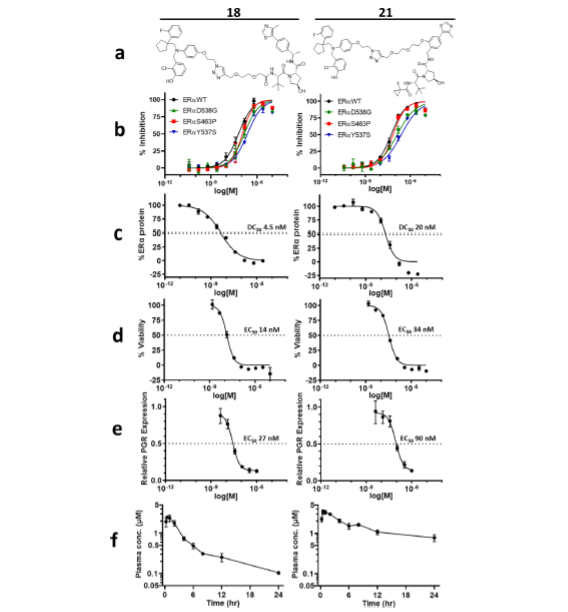
Figure 3. Biochemical, cellular, and mouse Pharmacokinetic (PK) properties of ERα PROTACs, 18 and 21. (a) Structures of 18 and 21. (b) Biochemical inhibition using purified recombinant ERα WT and mutants D538G, Y537Y, and S463P. (c) Degradation of ERα in MCF7 cells treated for 24 h followed by in-cell Western analysis of ERα protein normalized to tubulin. The graphs represent normalized ERα levels. (d) MCF7 viability assay with cells treated for 7 days followed by an end-point viability by CellTiter-Glo. (e) Downstream Progesterone Receptor (PGR) expression analysis in MCF7 cells treated for 24 h followed by RNA extraction, first strand synthesis, and qPCR analysis for progesterone receptor RNA estimation, normalized to TBP (Tata-binding protein); graphs represent normalized PGR levels. (f) Mean plasma concentration (μM) for 10 mpk (mg/kg) subcutaneous dosing in CD1 mouse (n = 3). Curves shown are from a single run with technical replicates.
Reprinted with permission from Disch J.S. et al., Journal of Medicinal Chemistry 2021 64 (8), 5049-5066, DOI: 10.1021/acs.jmedchem.1c00127. Copyright © (2021) American Chemical Society.
Testing Tumor Knockdown In Vivo
We compared the efficacies of compounds 18 and 21 with that of standard-of-care Fulvestrant in a wild type, estrogen-dependent MCF7 xenograft model (Fig. 4). Compound 21 decreased tumor volume almost as well as Fulvestrant. These results demonstrate that 21 is excellent for inhibition and degradation of ERα, both in vitro and in vivo. The ERα-binder of 21 was identified directly from the DEL screen with minimal optimization.
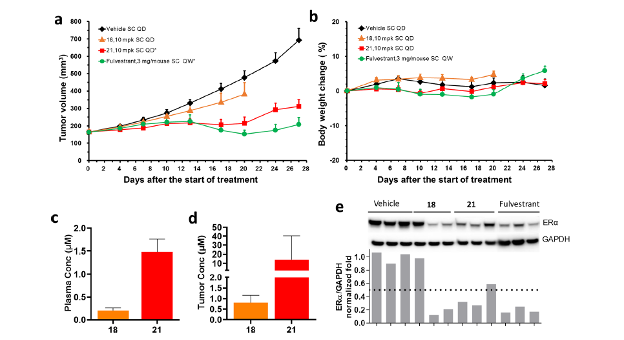
Figure 4. In vivo efficacy and Pharmacokinetic and Pharmacodynamic (PK/PD) study showing efficacy in MCF7 xenograft model. (a) Comparison of antitumor activity of 18 and 21 to Fulvestrant in WT Estrogen-dependent MCF7 xenograft model. Compounds 18 and 21 were administered subcutaneously (SC) once daily (QD) at 10 mg/kg. Fulvestrant was dosed SC once a week (QW) at 3 mg/mouse. Data represent the mean tumor volume ± SEM (n = 8). *p < 0.05 versus vehicle control (Student’s t-test). (b) All compounds were well tolerated with no changes in body weight. (c, d) Terminal Plasma PK and tumor PK of 18 and 21 at 3 h post-last dose (Day 20; 18) and (Day 27; 21). (e) PD analysis from tumor bearing mice treated with compounds mentioned above for 3 days. Western blot analysis was performed to demonstrate reduction in ERα levels upon treatment with 18, 21, and Fulvestra.
Reprinted with permission from Disch J.S. et.al., Journal of Medicinal Chemistry 2021 64 (8), 5049-5066, DOI: 10.1021/acs.jmedchem.1c00127. Copyright © (2021) American Chemical Society.
X-Chem Can Help You Identify and Assemble the Binders, Linkers, and E3 Ligands for Your Next PROTAC Project
This case demonstrates the potential and power of DEL for PROTAC applications. As the leader in DEL technology, X-Chem can help you screen DNA-encoded chemical libraries to identify novel binders as we did here and set you on the path to discovery of a groundbreaking compound. From hit identification for challenging targets to biochemical and biophysical profiling and elucidation of SAR through medicinal and synthetic chemistry, we’re here to assist you in your quest to unearth tomorrow’s life-saving therapies. Explore our website to learn more.
Reference
- Disch, J.S. et al. Bispecific Estrogen Receptor α Degraders Incorporating Novel Binders Identified Using DNA-Encoded Chemical Library Screening. Journal of Medicinal Chemistry 2021, 64 (8), 5049-5066.


