
One of the most attractive aspects of DNA-encoded library (DEL) technology is the vast number of compounds that can be generated and screened. DEL practitioners appreciate the speed and convenience of one-pot affinity selection, the ability to conduct parallel selections under various conditions and the depth of the selection output — all of these are equally powerful advantages. But for many, the entry point for their interest in DEL technology is the prospect of screening “billions and billions” of compounds. Since this aspect of the platform is so intuitively appealing, many DEL operators tout library size as part of their competitive advantage. Decision-makers in pharma and biotech often benchmark the library sizes of various providers as part of their assessments when selecting whose services to seek.
With so much attention on library size, we decided to ask a simple question: Are larger libraries actually better? X-Chem has accumulated enough data over the years that we should be able to directly address this question. To do so, we decided to look at our 100 licensed programs, which encompass over 700 distinct chemotypes, or “families,” in our nomenclature.
Why look at licensed families? Licensed families are those that we know had the potency, novelty and structural attractiveness to justify clients’ investment in follow-up chemistry. In essence, we are defining “better” in the question above as “better able to supply quality starting points that medicinal chemists want to pursue.” And really, what other metric counts? Why else do we screen if not to advance drug discovery?
A brief word about the use of the term “library”: At X-Chem, we use “library” as short-hand for “combinatorial library,” which is defined by Wikipedia as a “multi-component mixture[s] of small-molecule chemical compounds that are synthesized in a single stepwise process [italics added].” We have synthesized over 100 libraries at X-Chem, which can contain anywhere from 500,000 to 60 billion compounds. Since at X-Chem we rarely repeat a particular scheme, each library is unique and distinct from the others. Some may use the term “library” to refer to the entirety of their collection of synthetic mixtures, which may also be termed “sub-libraries.” Others may use the term to describe the product of a particular synthetic operation regardless of its uniqueness or redundancy. For our analysis, we will use the term in its classical sense.
To conduct our analysis, we determined which library each of our >700 licensed families arose from. We then attributed the libraries to half-log bins of library size and tallied up the number of licensed families in each bin. The results of this analysis are shown below.
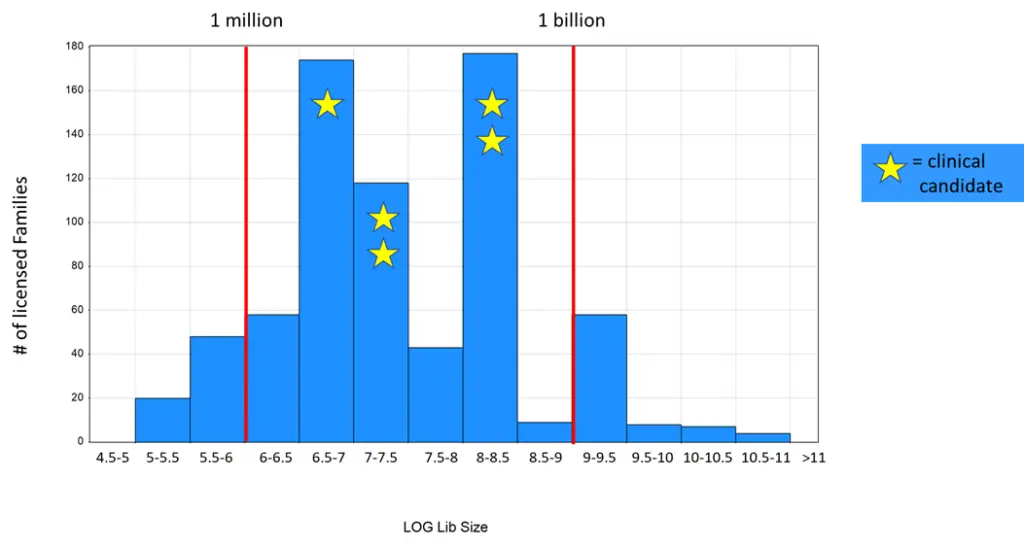
We can see that the most productive half-log bins are the 6.5-7, 7-7.5 and 8-8.5 bins. These correspond to library sizes in high single-digit million, low double-digit million and low triple-digit million ranges, respectively. There is no increase in productivity at larger library sizes. Interestingly, the 9-9.5 bin is equally productive as the 6-6.5 bin, despite being 1,000x bigger numerically (alternatively, one could say that the smaller bin is 1,000x more productive if productivity is defined as a function of compound count). We also examined where our five clinical candidates (two kinases, two PPIs and an enzyme) came from. We can see that these candidates all arose from those same three most productive bins.
Our interpretation of this analysis is that the answer to our question above is an emphatic “no.” We see no evidence that larger libraries are better, in that they do not provide a higher probability of yielding actionable starting points for medicinal chemistry (and eventually candidates). Rather, libraries with between three and 300 million compounds seem to represent a sweet spot in terms of library size. Put another way, we can see that ~90% of our licensed families come from libraries with <1 billion compounds, which comprise ~5% of our total compound count across all libraries. We are confident that this analysis is not confounded by library age, in that older libraries have undergone more screens and therefore have exaggerated productivity. If anything, such a skew would run in the other direction, since our largest libraries tend to be our older libraries and have seen more targets on average.
We believe that there is a role for large libraries, in the generation of tool compounds, for programs pursuing non-oral dosing and for particularly difficult targets. But overall, the trend is clear: smaller libraries drive drug discovery more effectively than large ones. At X-Chem, we feel that this analysis justifies our ongoing emphasis on chemical properties and atom economy. Because isn’t that why we do this, to find drugs that improve human health? That should be the ultimate criteria for any innovative technology in this industry.


